- Home
- Genomes
- Genome Browser
- Tools
- Mirrors
- Downloads
- My Data
- Projects
- Help
- About Us


Genome Browser
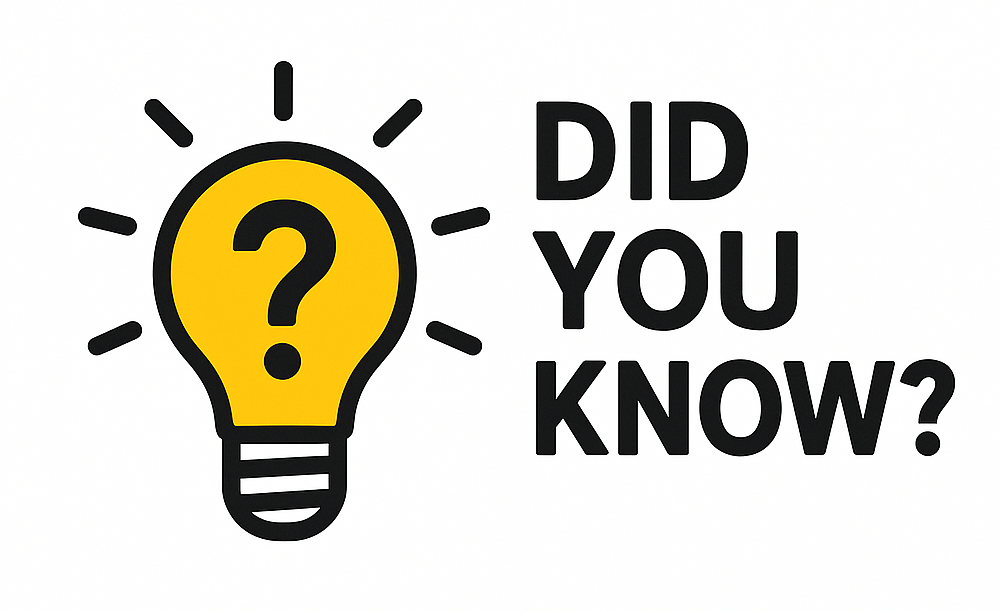

The Track Collection Builder (My Data > Track Collection Builder) allows multiple signal tracks to be copied and grouped together into one composite. Signal tracks in a composite container can then be overlaid, auto-scaled, sorted by similarity, and more.
Feel free to contact us if you are interested in attending a workshop, or meeting someone from the team to collaborate, get help, or ask any questions at the meetings. See our Poster Gallery.

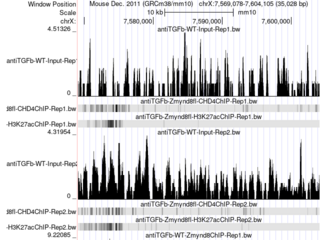
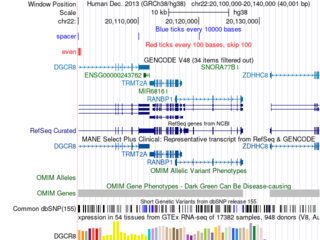
Click the images to explore the latest Public Sessions on our main site.